When you begin working on dinoflagellates, you quickly learn that they have (very) big genomes. But how big really?
Recently, I was preparing a presentation on dinoflagellates and I was faced with this question, so I investigated a bit. I found this great data visualisation by Tom White that plotted genome size in different groups of organisms. The data listed the size (in billions of base pairs, Gbp) of sequenced genomes on the NCBI database as of 2019. Because Mr. White was kind enough to publish the data he used and the code he wrote on his github, I had a look at it.
Genome size olympics
Among the 48 790 organisms listed in the data, I only found 4 dinoflagellates (I may have missed some). Because dinoflagellate genomes are notoriously big, and because they are quite strange, only a few genomes have been sequenced, and most of them very recently. Fortunately for me, Lin (2024) recently published a review article on dinoflagellate genomics. In his Table 1, he lists the published sequenced genomes of dinoflagellates and their size (in millions of bp, Mbp). How convenient for us!
So I made a little R script to visualize where these genomes fit in terms of size, compared to other organisms from all across the tree of life. Data and code are available on my github.
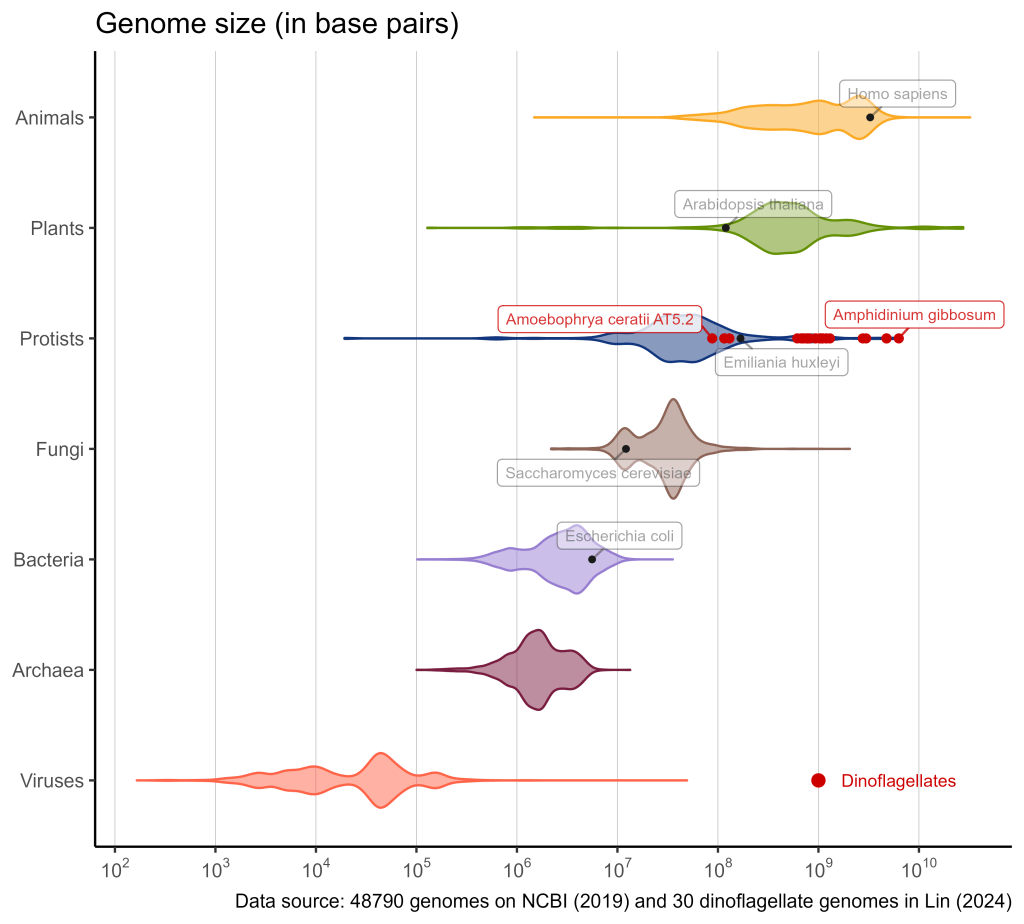
Apparently, the dinoflagellate genomes split in 3 groups of size.
- The smallest ones (close to 108 bp) are parasites such as Amoebophrya, who are close relatives of the core dinoflagellates (Dinophyceae). Funnily enough, some of them infect other dinoflagellates.
- The medium ones (around 109 bp) are mainly symbiotic taxa found within multicellular organisms, notably corals. They are often called Zooxanthellae and are probably among the most famous dinoflagellates.
- The last group (>2.5 x 109 bp) consists of 4 organisms : 2 strains of the polar dinoflagellate Polarella glacialis, Prorocentrum cordatum and Amphidinium gibbosum. All of them are free-living (i.e., neither symbionts nor parasites).
Parasitic and symbiotic organisms often have reduced genomes compared to their free-living relatives1. It also seems to be the case in dinoflagellates! The second and third dino groups appear to be the biggest known protist genomes. Although they are minute animalcules, free-living dinoflagellates surpass many plants and animals in terms of genome size!
But these are only the sequenced genomes we have for dinoflagellates. Some unsequenced ones are thought to be much, much bigger…
~225 Gbp ?!
Sequencing the nuclear DNA allows you to know the genome size of an organism. But there are other ways to estimate it. From what I read, it generally involves staining the DNA inside the cell with a fluorescent dye, measuring the amount of fluorescence and comparing it with calibration standards (cells the genome size of which is documented.)
Scientists who conducted such experiments on the dinoflagellate Prorocentrum micans have estimated genome sizes of up to 225 Gbp2. This is gigantic. It would certainly be one of the biggest known genomes, if not the biggest.
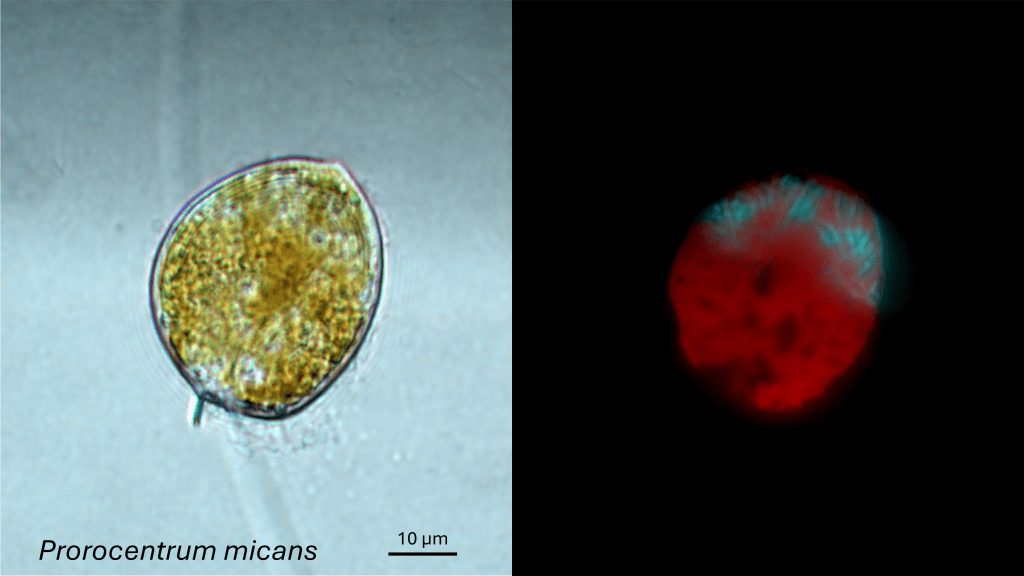
But it seems that such observations of “giant” genomes in dinos may be unreliable. As explained by Hidalgo et al. (2017) in their Box 1, several methodological uncertainties cast doubt on the reports of genomes > 100 Gbp in protists. They conclude: “Overall, although it is clear that the genomes in these eukaryotic lineages are large, only by estimating their sizes using best-practice techniques will we know just how big their genomes are compared with [the biggest known plant genomes].”
So, as far as I understand, the answer to the question in the title of this post could be: “very big for protists, quite big compared to many plants and animals, and potentially the biggest genomes in all living organisms but really we don’t know for sure”.
You may find this answer disappointing, but to me it’s very exciting, as it means there are still many things to be discovered about dinoflagellate genomes3.
But why are dinoflagellate genomes so big? This is a complicated topic that would deserve its own article, written by someone who actually knows things about genomics4. Hopefully I can persuade one of my esteemed geneticists colleagues to write on this blog one day!
This blog post and all the media it contains (figure, images) was produced by me, and is under a Creative Commons Attribution (CC BY) licence. You can reuse it freely as long as you cite the author (Victor Pochic) properly. I would like to sincerely thank Tom White for making his data and code public, and indirectly allowing me to do this work.
- Husnik et al. (2021) ; Nakayama et al. (2019) ↩︎
- LaJeunesse et al. (2005) ↩︎
- Actually, I know for a fact that the genomes of very interesting dinoflagellates are being sequenced as I’m writing this blog post. ↩︎
- Which is not my case, I just have a weird obsession with everything dinoflagellate :). But if you are interested in reading further on dinoflagellate genomics, you can start with Lin’s review (2024). ↩︎

Leave a comment